ANGIOGENES: A knowledge database for angiogenesis
Institute of Cardiovascular Regeneration, Goethe University Frankfurt



| Operator | Illustration | Explanation |
|---|---|---|
| and |  |
The "and" operation returns only elements common to both sets. |
| or |  |
The "or" operator returns only elements common to either or both sets. |
| not |  |
The "not" operator returns only elements not in the second set. |
|
This is a cheat-sheet for the advanced search.
A more comprehensive explanation can be found on the
help page
.
The advance search was designed in response to the high expression specificity of lncRNAs. We hope users will recognize its utility as queries are are only limited by ones imagination and the RAM memory of the server. | ||
| Expression | Example (Links run searchs) | Description |
|---|---|---|
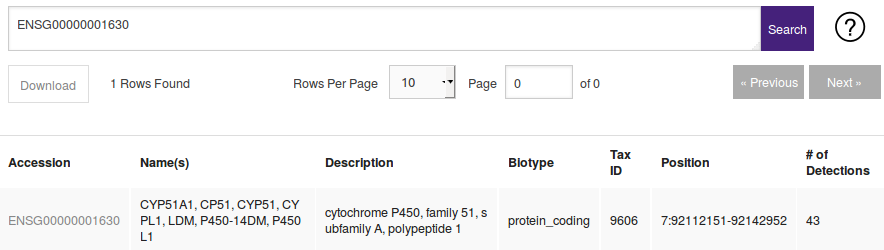
| No tag. | MALAT1 | When no tag is specified the query is treated as an accession or gene name. Wildcards are supported. |
| SPECIFIC:(condition) | SPECIFIC:"HAoEC--Heart--Normal" | Returns sequences specific (i.e. not expressed elsewhere.) to the specified condition. See "Supported Conditions" on the Help page for a list of searchable values. |
| ENRICHED:(condition) | ENRICHED:"HAoEC--Heart--Normal" | Returns sequences with higher than average expression in the given condition. See "Supported Conditions" on the Help page for a list of searchable values. |
| EXPRESSED:(condition) | EXPRESSED:"HAoEC--Heart--Normal" | Returns transcripts expressed in the given condition. See "Supported Conditions" on the Help page for a list of searchable values. |
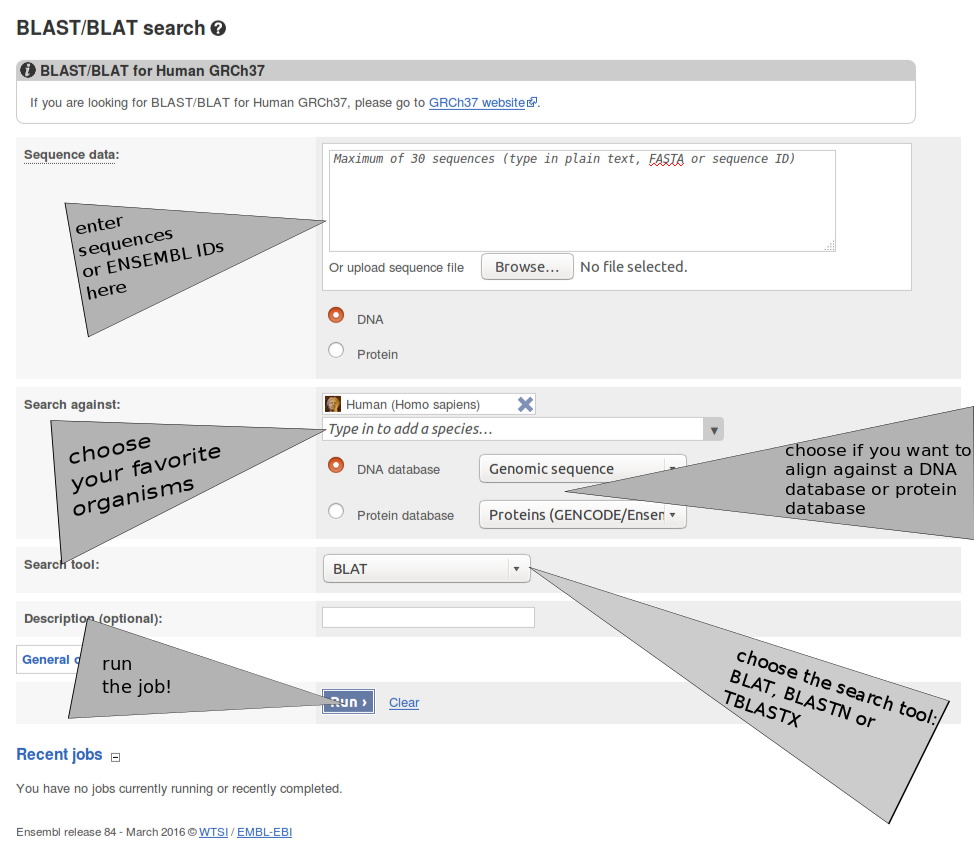
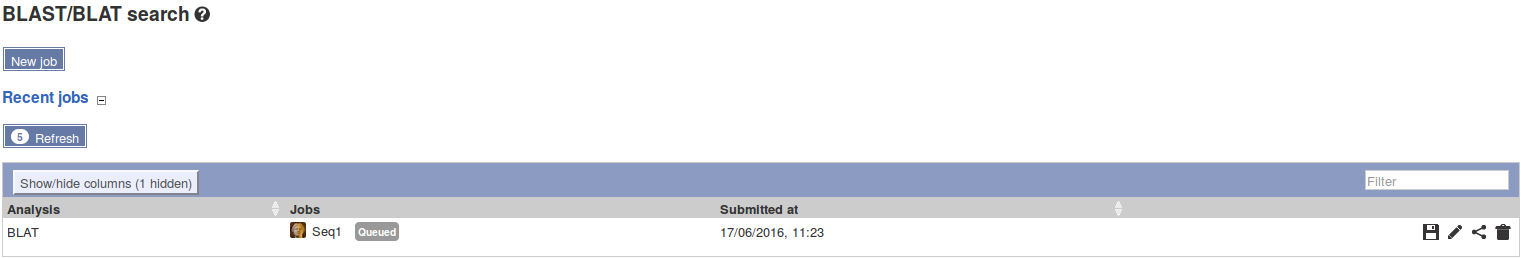
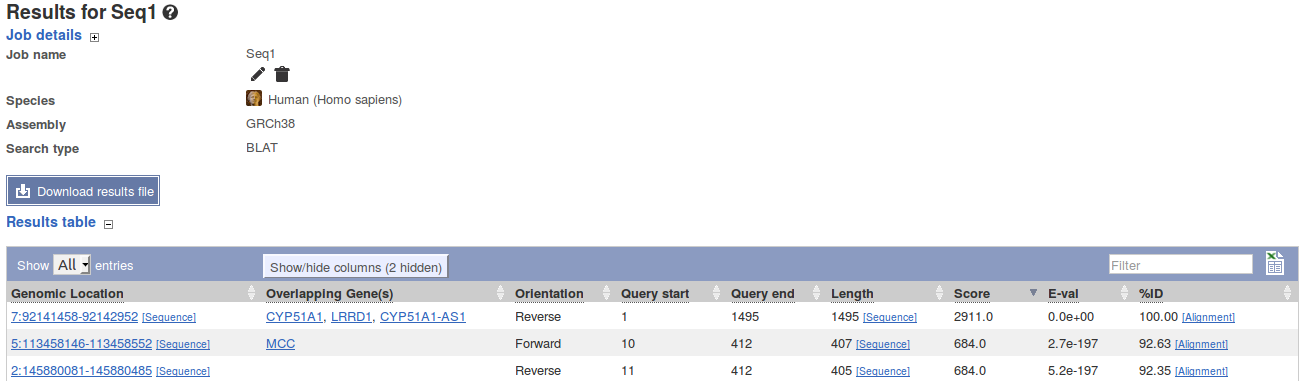
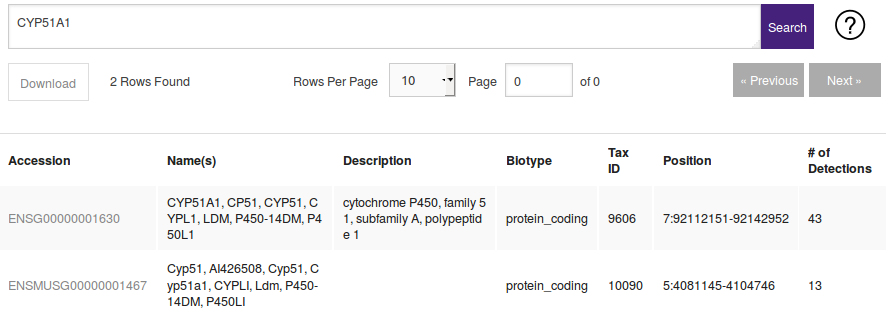
| HOMOLOGS:(Gene Accession) | HOMOLOGS:ENSG00000003436 | Returns homologs of searched genes. For a BLAST/BLAT search see Finding Homologs |
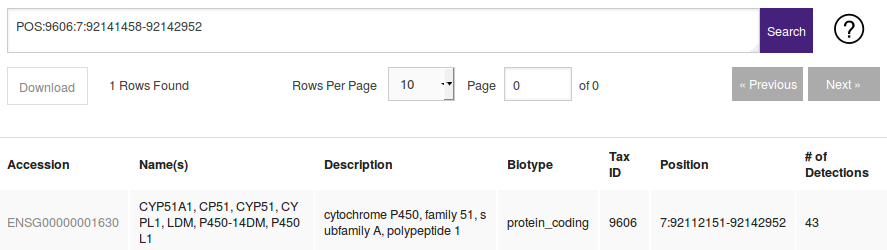
| POS:(tax_id):(chromosome):(start)-(end) | POS:9606:1:914880-969309 | Returns regions, genes, or transcripts falling withing the given position. |
| BIOTYPE:(biotype) | BIOTYPE:lincRNA | Returns regions, genes, or transcripts which contain a transcript annotated as the given biotype. |
| REGIONTYPE:(CGP|UCNEbase|VISTA) | REGIONTYPE:UCNEbase | Returns regions which contain the given region type. If searching for genes or transcripts, genes and transcripts contained within the given region type will be returned. For detail, please refer to C-It-Loci database: http://c-it-loci.uni-frankfurt.de |
| TAXID:(7955|9606|10090) | TAXID:7955 | Returns regions, genes, or transcripts associated with the given taxonomy. Taxonomy IDs (TAXID) are: 7955 (zebrafish), 9606 (human), and 10090 (mouse). |
| GO:(GO term accession) | GO:0005515 | Returns regions, genes, or transcripts which contain an annotation of the given Gene Ontology (GO) term. |
| Boolean Operators | ||
| and | TAXID:7955 and BIOTYPE:lincRNA | Returns accessions existing between two query sets. |
| or | BIOTYPE:lincRNA or BIOTYPE:antisense | Returns accessions existing in either of two query sets. |
| not | POS:9606:1:814880-3069309 and not POS:9606:1:1714880-2169309 | Returns accessions not in the query. |
| () | BIOTYPE:lincRNA and (TAXID:9606 or TAXID:10090) | Increases the precedence of a query. |
| * | gata* | Specifies where to use wildcards in query. Note: wildcards are only supported for tagless searches. |
| More Examples | ||
| Behavior | Query | Explination |
| Return all lncRNAs |
BIOTYPE:lincRNA or BIOTYPE:processed_transcript or BIOTYPE:3prime_overlapping_ncrna or BIOTYPE:antisense or BIOTYPE:non_coding or BIOTYPE:retained_intron or BIOTYPE:sense_intronic or BIOTYPE:sense_overlapping |
Using the "or" operator, link all sub-types of lncRNA. See Ensembl's documentation for a description of each biotype. |
| Return lincRNAs falling within Conserved Gene Pairs (CGPs) | BIOTYPE:lincRNA and REGIONTYPE:CGP | One lncRNA characteristic is a lack of sequence conservation. Therefore, by using conserved gene pairs or other highly conserved regions, one can quickly find syntetic regions to hunt for related lncRNAs. |
| TAXID | Supported Condition |
|---|---|
| 10090 | C166--Yolk sac--EPAS1 KD |
| 10090 | C166--Yolk sac--Normal |
| 10090 | Tie2plus EC--Cerebral cortex--Normal |
| 10090 | Tie2plus EC--kidney--Normal |
| 10090 | mES--Embryoid body--Gata1&2KO |
| 10090 | mES--Embryoid body--Normal |
| 10090 | mES--Embryoid body--SclKO |
| 9606 | BECs--Skin--Normal |
| 9606 | BECs--Skin--Under laminar flow |
| 9606 | HAoEC--Heart--Normal |
| 9606 | HAoEC--Heart--SAHA treatment |
| 9606 | HAoEC--Heart--TSA treatment |
| 9606 | HHSEC--Liver--Normal |
| 9606 | HMVEC-D--Skin--Normal |
| 9606 | HUVEC--Umbilical vein--EPAS1 KD |
| 9606 | HUVEC--Umbilical vein--Normal |
| 9606 | HUVEC--Umbilical vein--VEGF treatment 12h |
| 9606 | HUVEC--Umbilical vein--VEGF treatment 1h |
| 9606 | HUVEC--Umbilical vein--VEGF treatment 4h |
| 9606 | HUVEC--Umbilical vein--Virilizer |
| 9606 | HUVEC--Umbilical vein--WTAP |
| 9606 | HUVEC--Umbilical vein--cell polyAminus |
| 9606 | HUVEC--Umbilical vein--cell polyAplus |
| 9606 | HUVEC--Umbilical vein--cytosol polyAminus |
| 9606 | HUVEC--Umbilical vein--cytosol polyAplus |
| 9606 | HUVEC--Umbilical vein--nucleus polyAminus |
| 9606 | HUVEC--Umbilical vein--nucleus polyAplus |
| 9606 | HUVEC--Umbilical vein--siScr |
| 7955 | LECs--Lymphatic--FACS isolation of Kaede photconverted red ECs |
| 7955 | Unknown1--Pharyngeal arch--Normal GFPplus |
| 7955 | Unknown2--Pharyngeal arch--Normal GFPminus |